Proteomics & Metabolomics: Assessment Question Bank
This question bank covers key concepts in Proteomics and Metabolomics. Designed for undergraduate-level students.
- Highly quantitative and reproducible
- Minimal sample preparation
- Non-destructive and provides direct structural information
- Lower sensitivity compared to mass spectrometry (MS), limiting detection of low-abundance metabolites
(Select the single best answer)
Case-Based Questions (16–20)
A 55-year-old patient is being monitored for potential recurrence of colorectal cancer (CRC). Current imaging is inconclusive. A research team analyzes his annual blood plasma samples (from pre-diagnosis to now) using a multi-omics approach.
Part C: True or False (1 mark each)
[E] Answer True or False for the following statements.
Part C: Answer Key
Part D: Fill in the Blanks (1 mark each)
[E] Fill in the appropriate term in each blank.
Part D: Answer Key
Part E: Critical Thinking & Case Studies (5 marks each)
[H] Answer the following case studies with appropriate justification.
A) Metabolomics would provide the most direct functional insight because it directly measures the small-molecule substrates and products of energy metabolism (e.g., TCA cycle intermediates, lactate, ATP/ADP ratios), offering an immediate snapshot of mitochondrial dysfunction.
B) Both NMR or MS are acceptable with justification:
- NMR: Suitable for rapid, quantitative, and reproducible detection of major energy-related metabolites such as lactate, creatine, and amino acids in serum or urine.
- MS: Preferred when maximum sensitivity is required to detect low-abundance intermediates or to perform broad metabolite profiling from muscle biopsy samples.
A) Proteomic Strategy: Perform comparative proteomic or phosphoproteomic analysis on pre-treatment tumor biopsies from responders and non-responders. This could identify differential signaling pathways, drug targets, or resistance mechanisms associated with response.
B) Metabolomic Strategy: Analyze pre-dose and early-treatment plasma samples using MS or NMR to identify metabolic signatures (e.g., bile acids, lipid species, oxidative stress markers) that predict susceptibility to liver toxicity before clinical symptoms appear.
Grow both strains under controlled and drought-stress conditions. Collect leaf tissue at multiple time points and extract metabolites. Analyze samples using GC-MS and/or LC-MS to profile primary metabolites (sugars, amino acids) and secondary metabolites (antioxidants, stress hormones). Comparative analysis would identify metabolites such as proline, sugars, or antioxidants that accumulate preferentially in the drought-tolerant Strain B.
While genomics identifies underlying genetic variations, proteomics and metabolomics reveal the functional consequences and real-time physiological state of a biological system. Many diseases involve pathway dysregulation without direct genetic mutation, which can only be captured by studying proteins and metabolites.
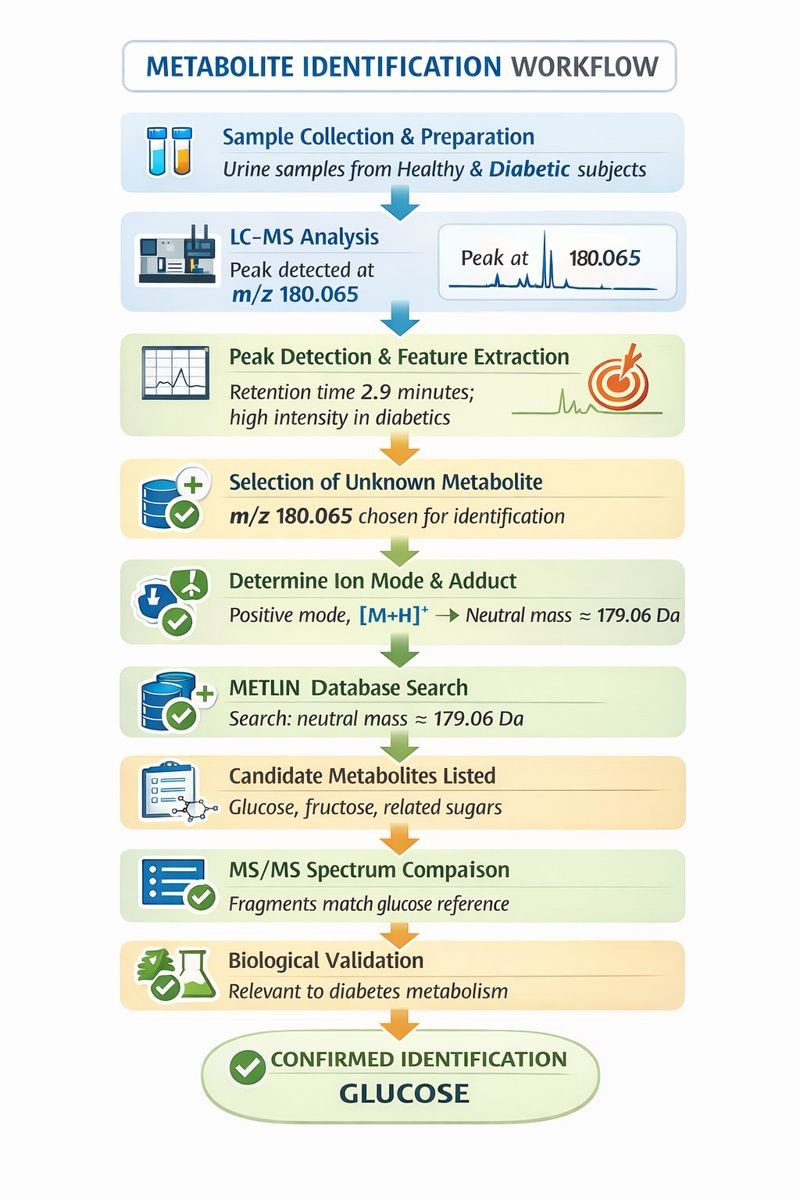
Flow Chart: METLIN Database–Based Metabolite Identification
Urine samples from healthy and diabetic subjects
Peak detected at m/z 180.065
Retention time: 2.9 minutes; high intensity in diabetics
m/z 180.065 chosen for identification
Positive mode, [M+H]+
Search neutral mass ~179.06 Da
Glucose, fructose, related sugars
Fragment ions match glucose reference spectrum
Relevant to diabetes-related metabolic pathways
Glucose
Questions on Human Metabolomics Database (HMDB)
Part A: Multiple Choice Questions
1. (E) What is the primary focus of the HMDB database?
- a) Human gene sequences
- b) Human protein structures
- c) Human small molecule metabolites
- d) Human disease symptoms
Answer: c
2. (E) Which search tool in HMDB is specifically designed for identifying metabolites using experimental data from a mass spectrometer?
- a) Text Query
- b) Sequence Search
- c) MS Search
- d) ChemQuery
Answer: MS Search
3. (H) Key feature distinguishing SMPDB from KEGG?
- a) It includes protein sequences.
- b) It is freely available.
- c) It integrates metabolic, drug, and disease pathways with a focus on small molecule visualization.
- d) It contains more pathways.
Answer: Integration of metabolic, drug, and disease pathways with small-molecule visualization
4. (E) Which is NOT part of the HMDB suite?
- a) DrugBank
- b) T3DB
- c) ProteinBank
- d) FooDB
Answer: ProteinBank
5. (H) If you have a pure, unknown compound and have collected its 1H NMR spectrum, which HMDB tool will give you the most direct and conclusive identification?
- a) Text Query with a guessed name
- b) 1D NMR Search with your peak list
- c) Browse by molecular weight
- d) Advanced Search by biofluid
Answer: 1D NMR Search with peak list
Fill in the Blanks
11. (E) The detailed metabolite information page is called a __________.
Answer: MetaboCard
12. (E) The database known for interactive pathway diagrams is __________.
Answer:SMPDB
13. (H) Best tool to find sulfur-containing metabolites linked to liver cancer?
Answer: Advanced Search
14. (E) Database containing pharmaceutical information?
Answer: DrugBank
15. (H) Drug–grapefruit interaction pathway examined in __________ database.
Answer: Drug metabolism pathway in SMPDB
Short Answer Questions
16. (E) Name two types of data in HMDB MetaboCard.
Answer: Chemical data, Clinical data
17. (E) Purpose of the Biospecimen table?
Shows normal and abnormal metabolite concentrations in biological samples.
18. (H) Advantage of Advanced Search?
Allows multi-parameter, relational queries for precise metabolite filtering.
19. (E) Database to find chemical components of apple?
Answer: FooDB
20. (H) Utility of SMPDB hyperlinks?
Enable direct navigation from pathways to detailed HMDB and DrugBank entries.
True or False
21. (E) HMDB includes endogenous and exogenous compounds.
Answer: True
22. (H) Sequence Search identifies unknown metabolites.
Answer: False
23. (E) FooDB is a recipe database.
Answer: False
24. (H) HML requires payment for digital spectra.
Answer: False
25. (E) SMPDB pathway maps are static.
Answer: False
Critical Thinking Skills
1. Novel Metabolite Identification in Kidney Disease
A metabolomics researcher identifies a novel compound in urine samples from patients with a rare kidney disease...
First, the MS Search tool in HMDB would be used with the exact mass (181.0739 Da) and a low ppm tolerance to generate candidate metabolites. The UV absorption at 260 nm suggests a conjugated or aromatic structure, helping further filter results. Next, the MetaboCards of top candidates would be examined for urine presence using the Biospecimen data and disease relevance via Clinical Data fields. SMPDB would then be used to identify involvement in kidney-related metabolic pathways. Finally, NMR Search or reference spectra comparison would confirm the metabolite’s identity.
2. HMDB Text Query vs Advanced Search
When would you choose HMDB's Text Query over Advanced Search?
Text Query is ideal for broad, exploratory searches using general names or concepts. For example, searching "ketone bodies" quickly retrieves acetoacetate, beta-hydroxybutyrate, and acetone. Advanced Search is used for precise, multi-parameter queries, such as identifying metabolites within a specific mass range that are elevated in Alzheimer’s disease and present at low concentrations in CSF.
3. Value of HMDB–SMPDB Integration
Explain how HMDB and SMPDB together provide greater research value.
HMDB provides chemical identity, concentration ranges, and disease associations for metabolites such as succinate. SMPDB adds biological context by visualizing the metabolite within pathways like the TCA cycle. Integrated navigation allows researchers to link biochemical abnormalities directly to disease mechanisms, such as mitochondrial dysfunction in heart failure.
4. Investigating Food Contamination Using HMDB Suite
How could FooDB and T3DB be used in public health investigations?
A researcher could identify suspected toxins using T3DB based on symptoms or chemical class. Natural sources and contamination routes listed in T3DB could then be cross-referenced with FooDB to identify contaminated foods. Links to HMDB and SMPDB would help assess human metabolic impact and toxicity pathways.
5. Defending the Importance of HMDB
Why is HMDB still important despite AI-based metabolite prediction?
HMDB provides curated experimental data that AI predictions cannot replace. This includes real MS/MS and NMR spectra for compound validation, measured clinical concentration ranges in human samples, and manually curated disease associations from peer-reviewed studies. These data are essential for experimental confirmation and biomedical discovery.